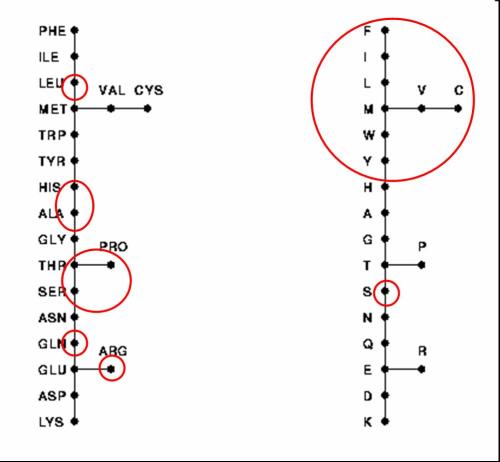
It has been argued that no engineer would have used cytosine as part of the genetic material because of its predisposition for deamination. But it’s exactly this predisposition that might cause an engineer of evolution to include it.
Life itself appears to have been designed to minimize errors. The universal nature of the proof-reading/repair machinery, the optimized genetic code, and the G/C:A/T parity code all converge on this point. Yet despite this design logic, there is the interesting fact that cytosine is especially prone to deamination, where the removal of its exocyclic amino group converts it into uracil (a base normally found in RNA). Uracil does not exist in DNA, thus it can be effectively detected and removed by repair enzymes. However, if not detected and repaired, it can base pair with adenine, meaning that it would specify adenine during DNA replication. In a subsequent round of replication, the adenine in turn would specify thymine. The bottom line is that spontaneous deamination of cytosine can lead to a base substitution known as a transition, where C is replaced by T (and G is replaced by A on the other strand of DNA). We might expect such mutations to be quite common, as the rate constant for cytosine deamination at 37 degree C in single stranded DNA translates into a half-life for any specific cytosine of about 200 years. In fact, such high rates of deamination led researchers Poole et. al to complain of “confounded cytosine!” [1]
We would thus seem to have two contradictory lines of evidence. On one hand, there is the growing list of evidence to support the hypothesis that error correction was an important principle guiding the design of life. Yet the incorporation of cytosine works against such efforts, given its predisposition to spark a mutation. In fact, Poole et al. go so far as to argue, “Any engineer would have replaced cytosine, but evolution is a tinkerer not an engineer.” From a design perspective, how might these contrary dynamics be reconciled? That is, given the emphasis on error correction, why would an engineer include cytosine?
Continue reading →